New approach to share clinico-genomic data
Researchers from UPM have developed a new tool that provides standard and shared clinico-genomic data
The Biomedical informatics group from Universidad Politécnica de Madrid has developed a Semantic Interoperability Layer (SIL) that uses standard terminologies as a vehicle for addressing two main challenges in multi-centric interoperability: harmonizing heterogeneities from different data sources as well as for integrating omic and clinical data to enhance prevention, diagnosis and treatments of diverse diseases such as breast cancer.
The introduction of omics data (from genomics, transcriptomics, proteomics and etc) involved in clinical treatment has led to a broad range of approaches to represent clinical information. Within this context, patient stratification across health institutions by omic profiling presents a complex scenario to carry out multi-center clinical trials. In the past, the patients needed to carry out a clinical study were found in the same hospital or healthcare center, however nowadays to recruit a minimum cohort of patients for a clinical study often involve multiple clinical institutions from different regions or countries.
 In this way, the data exchange among different centers is complex not only due to legal aspects but also technical issues. The information required for clinical studies is stored in each hospital and even in each department within a hospital in heterogeneous information systems that present different format and are encoded in diverse medical terminologies and languages.
In this way, the data exchange among different centers is complex not only due to legal aspects but also technical issues. The information required for clinical studies is stored in each hospital and even in each department within a hospital in heterogeneous information systems that present different format and are encoded in diverse medical terminologies and languages.
The Biomedical informatics group (GIB) from Universidad Politécnica de Madrid has been involved for the last years in the development of integration of clinico-genomic data from heterogeneous sources. In order to achieve a homogeneous access to the clinico-genomic data, researchers have developed a Semantic Interoperability Layer (SIL), a software layer based on terminologies and biomedical standards able to provide a homogeneous access to data.
The researcher Rául Alonso, a member from GIB, explains what SIL consists of. This tool is composed by four main elements: first, a Common Data Model, which is able to link in a standard way with the hospital information systems; second, a Core Dataset to codify the information and data from different hospitals; third, a Terminology binding between the common data model and the core dataset; and four, a set of services to access and manage the stored information”.
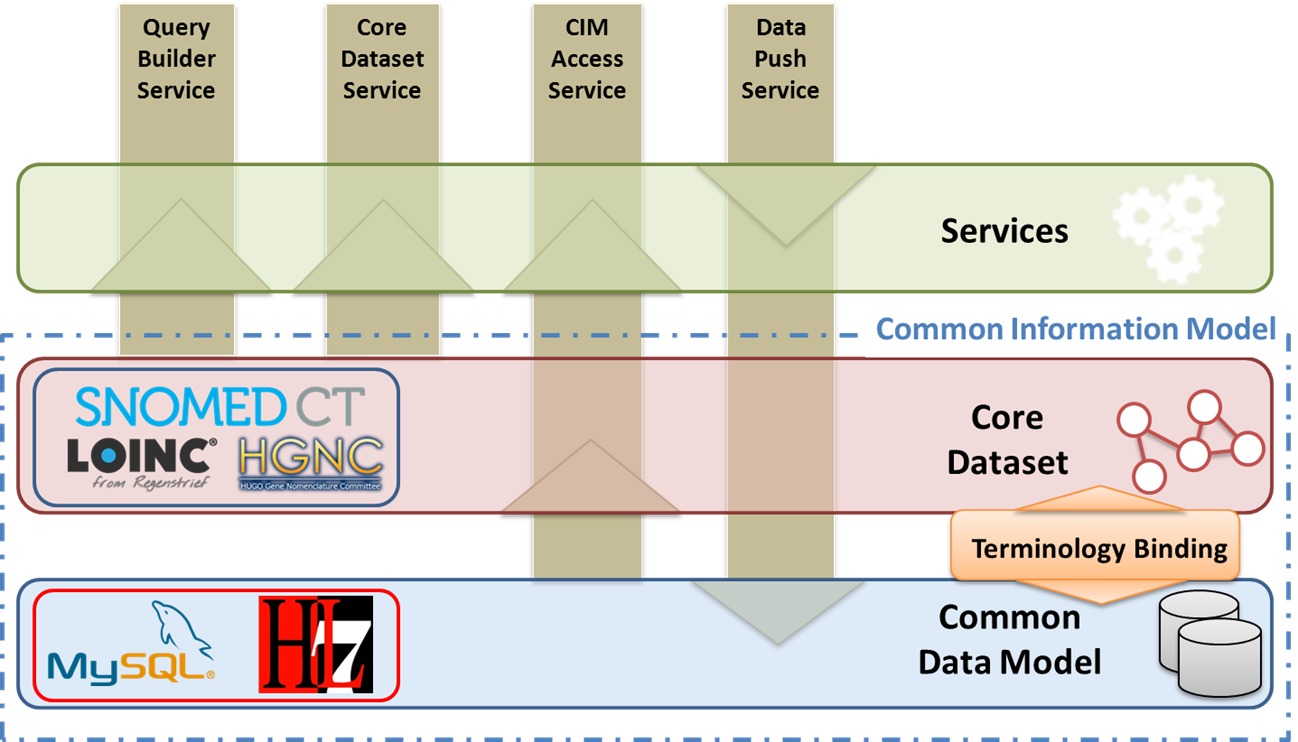
Elements of the Interoperability Layer
The new approach of this Semantic Interoperability Layer is that data is not only translated into the common vocabulary but is also standardized. For example, when there is a diagnosis of “neoplasia of the respiratory tract” in a hospital, such data is stored in a way comparable to other more specific or similar diagnosis from other hospitals, such as “primary adenocarcinoma of the lower lobe of the right lung” or “malignant tumor of the bronchi”.
Additionally, the Semantic Interoperability Layer is able to store genetic testing data in the same storage than clinical data in a way that the information can be homogenously stored and consulted.
This Semantic Interoperability Layer has been proved with real data from hospitals inSpain, Germany, Belgium, Holland and United Kingdom within the collaboration of several national and international projects. This work, carried out by members of the Biomedical informatics group (GIB) from Universidad Politécnica de Madrid at School of Computer Engineering, has been published in diverse international scientific journal.
Raúl Alonso Calvo, Sergio Paraíso Medina, David Pérez-Rey a, Enrique Alonso Oset, Ruud van Stiphout, Sheng Yu, Marian Taylor, Francesca Buffa, Carlos Fernández-Lozano. A semantic interoperability approach to support integration of gene expression and clinical data in breast cancer. Computers in Biology and Medicine 77: 179-186. August 2017


